Welcome to my professional website.
👋 My research interests concern vector-borne diseases. I aim to use sequencing methods and molecular strategies to develop cost-effective methods to detect arthropod-borne pathogens in the environment, or to assess vector population composition, densities and dispersion. I am open to explore within and beyond these frontiers through collaborations 😃 🚀
I inform readers that the views, thoughts, and opinions expressed in this site (if any) belong solely to the author, and not necessarily to the author’s employer, organization, institution or other group or individual. The opposite is also true, any opinions expressed my members of institutions I am involved with are not necessarily mine.
Download my resumé.
- Vector-borne diseases
- Mosquito-borne pathogens surveillance
- Interaction dynamics between mosquito-borne pathogens and their hosts
- Quantitative genetics applied to mosquito-viruses interactions
- Data visualisation
-
PhD (Hons) in Infectious diseases, 2010
Aix-Marseille II University (Marseille, France)
-
MSc in Human Pathology Infectious diseases, 2006
Aix-Marseille II University (Marseille, France)
-
BSc (Hons) in biological sciences, 2005
Salford University Greater Manchester, UK
Skills
100%
100%. I dip my toasts in my coffee and I can eat a frog or a snail…alive yes!
Too much!
50%
50%
As much as I can :)
Experience
Recent Posts
Projects
Featured Publications
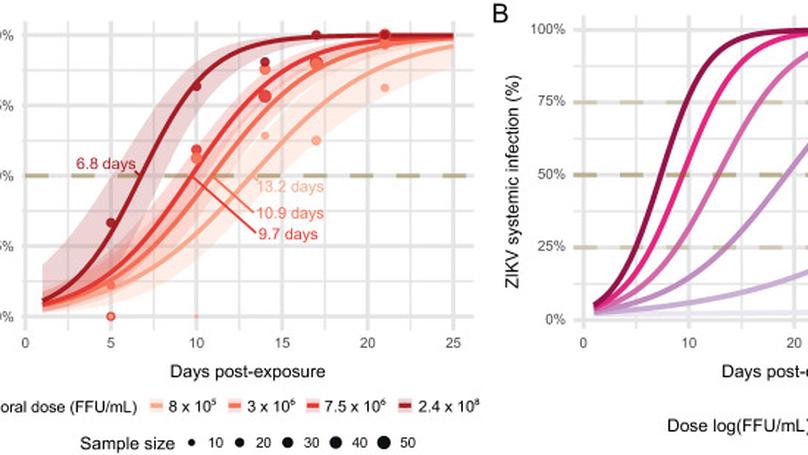
To assess Ae. albopictus vector potential for ZIKV and identify key virus outbreak predictors, we built a complete framework using the complementary combination of (i) dose-dependent experimental Ae. albopictus exposure to ZIKV followed by time-dependent assessment of infection and systemic infection rates, (ii) modeling of intra-human ZIKV viremia dynamics, and (iii) in silico epidemiological simulations using an Agent-Based Model. Our results reveal a low but existing epidemic potential of Ae. albopictus for ZIKV, that might explain the absence of large scale ZIKV epidemics so far in territories occupied only by Ae. albopictus. They nevertheless support active surveillance and eradication programs in these territories to maintain the risk of emergence to a low level.
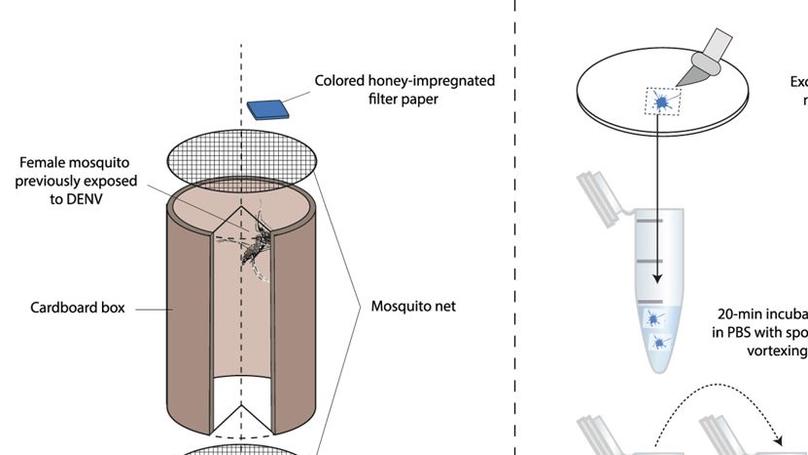
We report that individual Aedes aegypti mosquitoes release large amounts of dengue virus (DENV) RNA in their excreta that can be non-sacrificially detected over time following oral virus exposure. Further, we demonstrate that detection of DENV RNA in excreta from individual mosquitoes is correlated to systemic viral dissemination.
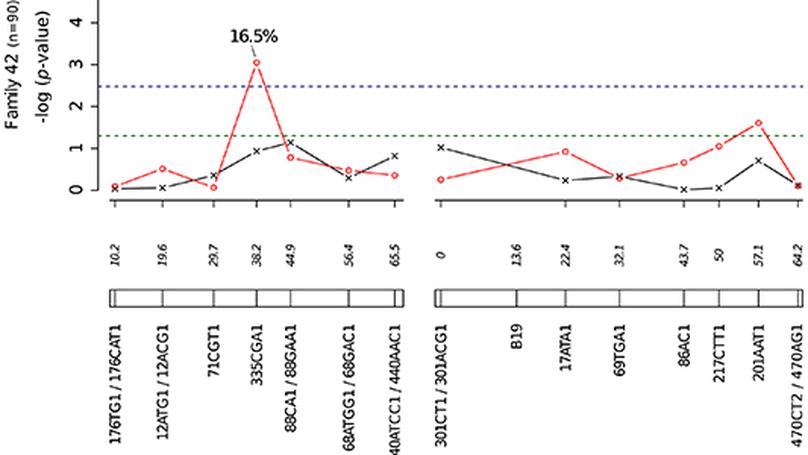
We developed an innovative genetic mapping strategy to survey G×G interactions using outbred mosquito families that were experimentally exposed to genetically distinct isolates of two dengue virus serotypes derived from human patients. Genetic loci associated with vector competence indices were detected in multiple regions of the mosquito genome. Importantly, correlation between genotype and phenotype was virus isolate-specific at several of these loci, indicating G×G interactions.
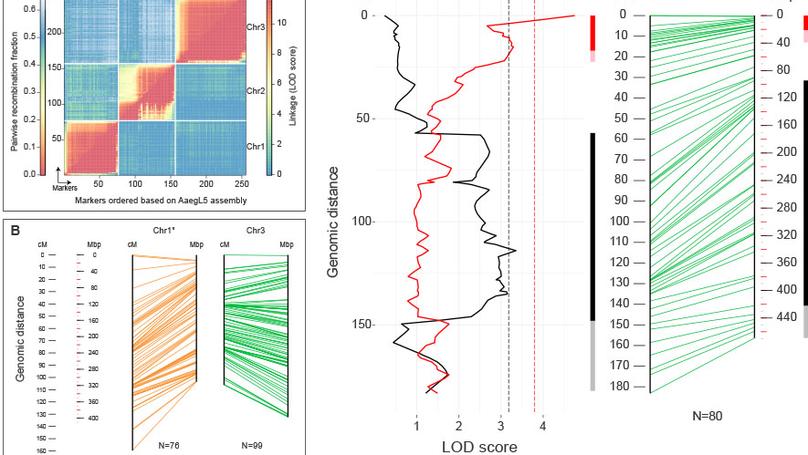
We participated to this study by illustrating the value of the new assembly (AaegL5) for mapping quantitative trait loci (QTLs). We used restriction site-associated DNA (RAD) markers to locate QTLs underlying dengue virus (DENV) vector competence.
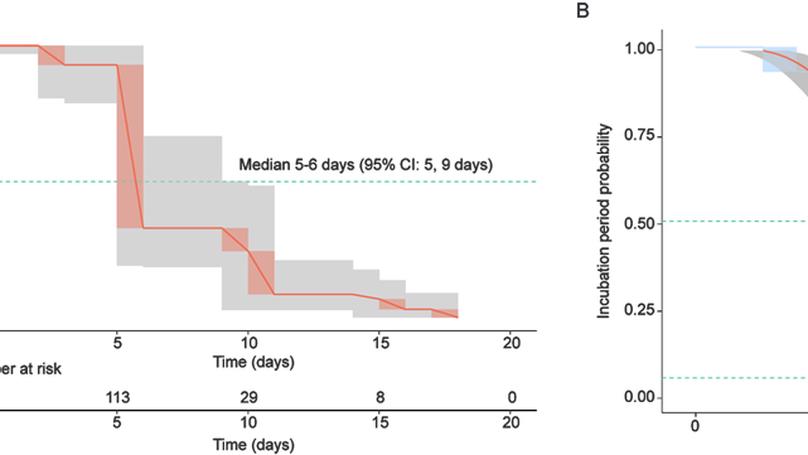
In this study, we estimated ZIKV incubation period distribution using time-to-event models adapted to interval-censored data based on declared date of travels from 123 symptomatic travelers returning from areas with active ZIKV transmission.
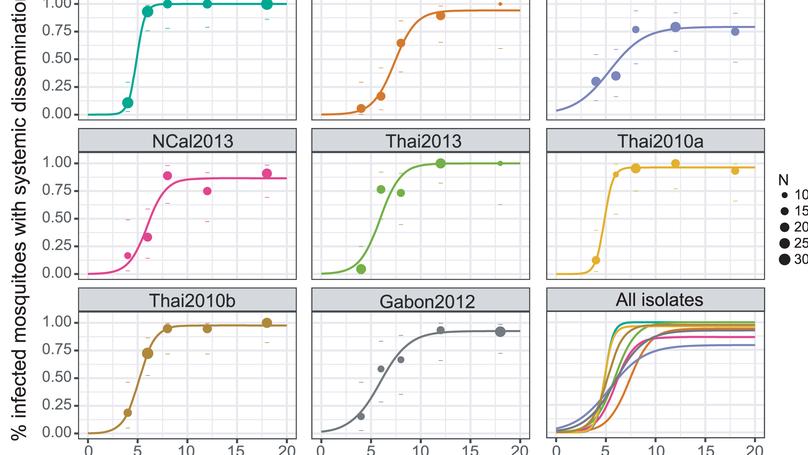
We combined newly generated empirical measurements in vivo and outbreak simulations in silico to assess the epidemiological significance of genetic variation in dengue virus (DENV) transmission kinetics by Aedes aegypti mosquitoes. We used a logistic model with three-parameters to estimate intra-mosquito virus dynamics based on the cumulative proportion of mosquitoes experimentally exposed to DENV with a systemic (disseminated) infection over time.
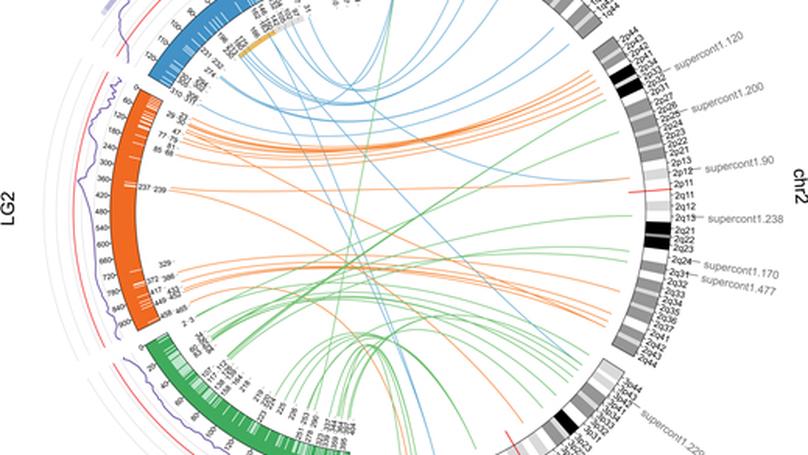
We provide evidence that sex chromosomes in Ae. aegypti are genetically differentiated between males and females over a region much larger than the sex-determining region (SDR). Our linkage mapping intercrosses failed to detect recombination between X and Y chromosomes over a 123-Mbp region (40% of their physical length) containing the SDR. Sex-differentiated genomic region was associated with a significant excess of male-to-female heterozygosity and contained a small cluster of loci consistent with Y-specific null alleles.
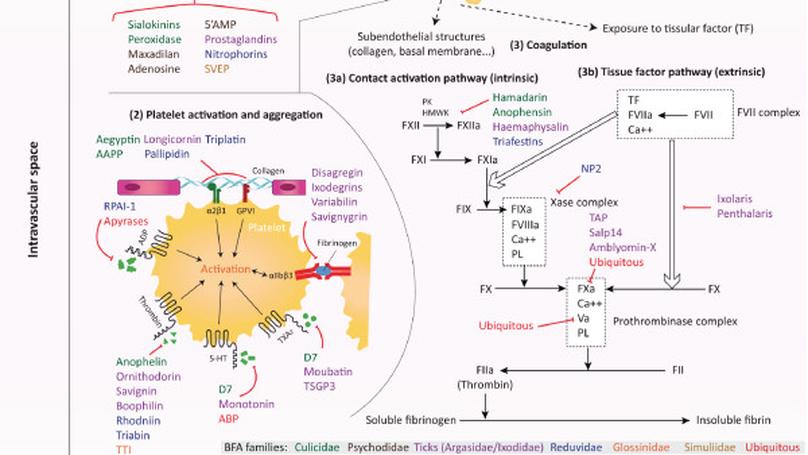
This paper provides a comprehensive overview of the pharmacological activity and immunogenic properties of the main salivary proteins characterised in various haematophagous arthropod species. The potential biological and epidemiological applications of these immunogenic salivary molecules are discussed with an emphasis on their use as biomarkers of exposure to haematophagous arthropod bites or vaccine candidates that are liable to improve host protection against vector-borne diseases.









